Quantitative Genomics of 30 Complex Phenotypes in Wagyu x Angus F1 Progeny
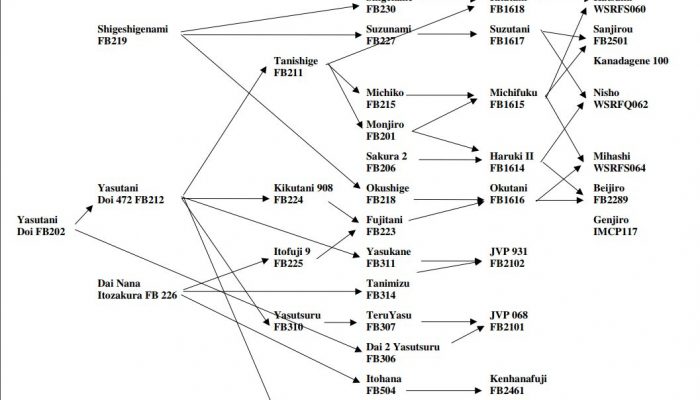
ABSTRACT: In the present study, a total of 91 genes involved in various pathways were investigated for their associations with six carcass traits and twenty-four fatty acid composition phenotypes in a Wagyu×Angus reference population, including 43 Wagyu bulls and their potential 791 F1 progeny. Of the 182 SNPs evaluated, 102 SNPs that were in Hardy-Weinberg equilibrium with minor allele frequencies (MAF>0.15) were selected for parentage assignment and association studies with these quantitative traits. The parentage assignment revealed that 40 of 43 Wagyu sires produced over 96.71% of the calves in the population. Linkage disequilibrium analysis identified 75 of 102 SNPs derived from 54 genes as tagged SNPs. After Bonferroni correction, single-marker analysis revealed a total of 113 significant associations between 44 genes and 29 phenotypes (adjusted P<0.05). Multiple-marker analysis confirmed single-gene associations for 10 traits, but revealed two-gene networks for 9 traits and three-gene networks for 8 traits.
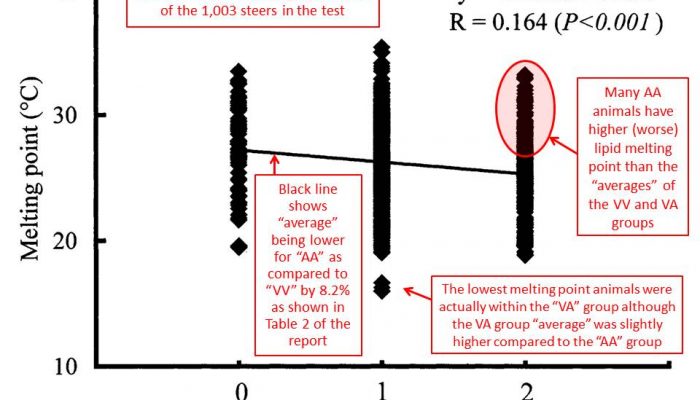
Particularly, we observed that TNF (tumor necrosis factor) gene is significantly associated with both beef marbling score (P=0.0016) and palmitic acid (C16:0) (P=0.0043), RCAN1 (regulator of calcineurin 1) with rib-eye area (P=0.0103), ASB3 (ankyrin repeat and SOCS box-containing 3) with backfat (P=0.0392), ABCA1(ATP-binding cassette A1) with both palmitic acid (C16:0) (P=0.0025) and oleic acid (C18:1n9) (P=0.0114), SLC27A1(solute carrier family 27 A1) with oleic acid (C18:1n9) (P=0.0155), CRH (corticotropin releasing hormone) with both linolenic acid (OMEGA-3) (P=0.0200) and OMEGA 6:3 RATIO (P=0.0054), SLC27A2 (solute carrier family 27 A2) with both linoleic acid (OMEGA-6) (P=0.0121) and FAT (P=0.0333), GNG3 (guanine nucleotide binding protein gamma 3 with desaturase 9 (P=0.0115), and EFEMP1 (EGF containing fibulin-like extracellular matrix protein 1), PLTP (phospholipid transfer protein) and DSEL (dermatan sulfate epimerase-like) with conjugated linoleic acid (P=0.0042-0.0044), respectively, in the Wagyu x Angus F1 population. In addition, we observed an interesting phenomenon that crossbreeding of different breeds might change gene actions to dominant and overdominant modes, thus explaining the origin of heterosis. The present study confirmed that these important families or pathway-based genes are useful targets for improving meat quality traits and healthful beef products in cattle.
CLICK HERE for the full research paper published online in the International Journal Of Biological Sciences.